
Machine Learning for Weed Detection (Video)
Video about the project EWIS: Evaluation and development of modern methods of artificial intelligence for automatic weed detection in sorghum using drones

Video about the project EWIS: Evaluation and development of modern methods of artificial intelligence for automatic weed detection in sorghum using drones
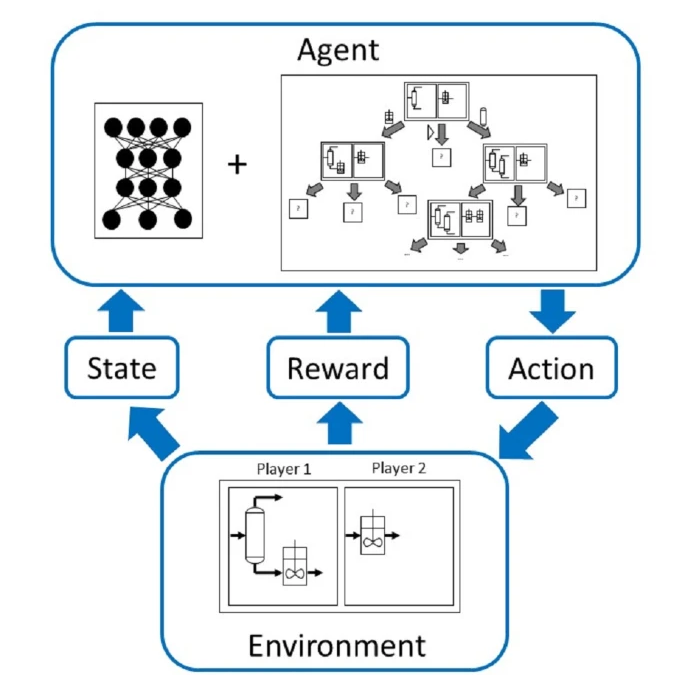
Automated flowsheet synthesis is an important field in computer-aided process engineering. The present work demonstrates how reinforcement learning can be used for automated flowsheet synthesis without any heuristics or prior knowledge of conceptual design. The environment consists of a steady-state flowsheet simulator that contains all physical knowledge. An agent is trained to take discrete actions and sequentially build up flowsheets that solve a given process problem. A novel method named SynGameZero is developed to ensure good exploration schemes in the complex problem. Therein, flowsheet synthesis is modelled as a game of two competing players. The agent plays this game against itself during training and consists of an artificial neural network and a tree search for forward planning. The method is applied successfully to a reaction-distillation process in a quaternary system.

In this chapter, we introduce the concept of RNA-Seq analyses. First, we start to provide an overview of a typical RNA-Seq experiment that includes extraction of sample RNA, enrichment, and cDNA library preparation. Next, we review tools for quality control and data pre-processing followed by a standard workflow to perform RNA-Seq analyses. For this purpose, we discuss two common RNA-Seq strategies, that is a reference-based alignment and a de novo assembly approach. We learn how to do basic downstream analyses of RNA-Seq data, including quantification of expressed genes, differential gene expression (DE) between different groups as well as functional gene analysis. Eventually, we provide a best-practice example for a reference-based RNA-Seq analysis from beginning to end, including all necessary tools and steps on GitHub: https://github.com/grimmlab/BookChapter-RNA-Seq-Analyses.

New publication about “Accurate machine learning-based germination detection, prediction and quality assessment of three grain crops” just got published in Plant Methods.
Our proposed machine learning-based method can help to speed up the assessment of seed germination experiments for different seed cultivars. It has lower error rates and a higher performance compared to conventional and manual methods, leading to more accurate germination indices and quality assessments of seeds.

New publication about “Network-guided search for genetic heterogeneity between gene pairs” just got published in Bioinformatics: https://academic.oup.com/bioinformatics/advance-article/doi/10.1093/bioinformatics/btaa581/5861532
We propose a novel method for finding pairs of interacting genes that are, upon combination, associated with a phenotype of interest under a model of genetic heterogeneity. We guide the interaction search using biological prior knowledge in the form of protein-protein interaction networks. Our method controls type I error and outperforms state-of-the-art methods with respect to statistical power. Additionally, we find novel associations for multiple A. thaliana phenotypes, and for a study of rare variants in migraine patients.